I am going to try to illustrate the mistake I mean exists in the legacy code with a small example:
from dolfin import *
class OutwardNormal(UserExpression):
def __init__(self, mesh, *arg, **kwargs):
super().__init__(*arg, **kwargs)
self.mesh = mesh
def eval_cell(self, values, x, cell):
c = Cell(self.mesh, cell.index)
normal = c.cell_normal()
scale = 1 if c.orientation == 0 else -1
values[0] = normal[0] * scale
values[1] = normal[1] * scale
values[2] = normal[2] * scale
def value_shape(self):
return (3,)
use_gmsh = False
if use_gmsh:
import meshio
import gmsh
order = 1
res = 0.25
gmsh.initialize()
outer_sphere = gmsh.model.occ.addSphere(0, 0, 0, 1)
gmsh.model.occ.synchronize()
boundary = gmsh.model.getBoundary([(3, outer_sphere)], recursive=False)
gmsh.model.addPhysicalGroup(boundary[0][0], [boundary[0][1]], tag=1)
gmsh.option.setNumber("Mesh.CharacteristicLengthFactor", res)
gmsh.model.mesh.generate(2)
gmsh.model.mesh.setOrder(order)
gmsh.write("gmsh_sphere.msh")
in_mesh = meshio.read("gmsh_sphere.msh")
meshio.write("gmsh_sphere.xdmf", in_mesh)
mesh = Mesh()
with XDMFFile("gmsh_sphere.xdmf") as f:
f.read(mesh)
else:
mesh = Mesh("../mixed-poisson/hdiv-l2/meshes/sphere_ico4.xml")
global_normal = Expression(("x[0]", "x[1]", "x[2]"), degree=1)
mesh.init_cell_orientations(global_normal)
n = OutwardNormal(mesh)
c_n = CellNormal(mesh)
x = SpatialCoordinate(mesh)
n_x = x / sqrt(dot(x, x))
class Projector:
def __init__(self, V):
u = TrialFunction(V)
v = TestFunction(V)
a = inner(u, v) * dx
self.A = assemble(a)
self.solver = LUSolver(self.A)
self.V = V
def __call__(self, f):
v = TestFunction(self.V)
b = assemble(inner(f, v) * dx)
u = Function(self.V)
self.solver.solve(u.vector(), b)
return u
Q = VectorFunctionSpace(mesh, "DG", 0)
projector = Projector(Q)
outward_n = projector(n)
cell_normal = projector(c_n)
spatial_normal = projector(n_x)
outward_n.rename("outward_normal", "outward_normal")
cell_normal.rename("cell_normal", "cell_normal")
spatial_normal.rename("spatial_normal", "spatial_normal")
with XDMFFile("normals.xdmf") as f:
f.write(outward_n, 0)
f.write(cell_normal, 0)
f.write(spatial_normal, 0)
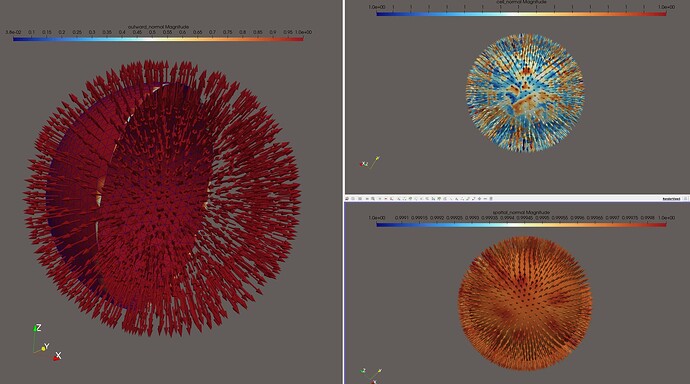
One can inspect all these normals in Paraview.
Using the mesh from the paper, I get:
Note that here, the “outward” normal is the only one that flips signs every now and then.
Next, if we use the gmsh mesh, we get:
where you see the exact same, fluctuating behavior in the “OutwardNormal”.
This means that it is very unclear to me how OutwardNormal should emulate the k from the paper, which is stated as:
Thus, when you look at the DOLFINx code, one should use an outward oriented normal, i.e.
k = global_orientation * ufl.CellNormal(mesh)
instead of ufl.CellNormal(mesh),
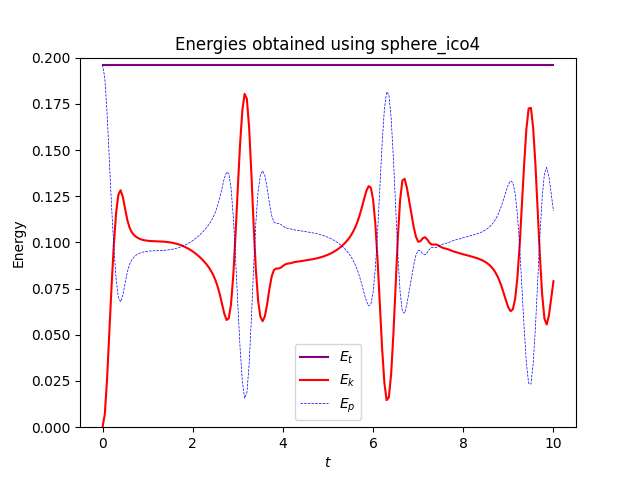
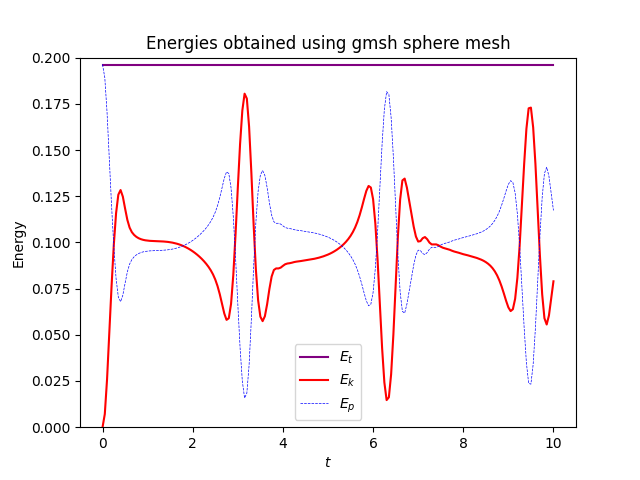
one gets very the exact same results for the old meshes and gmsh:
Full script is available below:
"""Linear shallow water equation on a manifold
Author: Jørgen S. Dokken
"""
from mpi4py import MPI
import numpy as np
import ufl
from typing import Optional
import basix.ufl
import dolfinx as df
from petsc4py import PETSc
from dolfinx.fem import petsc
class Projector:
"""
Projector for a given function.
Solves Ax=b, where
.. highlight:: python
.. code-block:: python
u, v = ufl.TrialFunction(Space), ufl.TestFunction(space)
dx = ufl.Measure("dx", metadata=metadata)
A = inner(u, v) * dx
b = inner(function, v) * dx(metadata=metadata)
Args:
function: UFL expression of function to project
space: Space to project function into
petsc_options: Options to pass to PETSc
jit_options: Options to pass to just in time compiler
form_compiler_options: Options to pass to the form compiler
metadata: Data to pass to the integration measure
"""
_A: PETSc.Mat # The mass matrix
_b: PETSc.Vec # The rhs vector
_lhs: df.fem.Form # The compiled form for the mass matrix
_ksp: PETSc.KSP # The PETSc solver
_x: df.fem.Function # The solution vector
_dx: ufl.Measure # Integration measure
def __init__(
self,
space: df.fem.FunctionSpace,
petsc_options: Optional[dict] = None,
jit_options: Optional[dict] = None,
form_compiler_options: Optional[dict] = None,
metadata: Optional[dict] = None,
):
petsc_options = {} if petsc_options is None else petsc_options
jit_options = {} if jit_options is None else jit_options
form_compiler_options = (
{} if form_compiler_options is None else form_compiler_options
)
# Assemble projection matrix once
u = ufl.TrialFunction(space)
v = ufl.TestFunction(space)
self._dx = ufl.Measure("dx", domain=space.mesh, metadata=metadata)
a = ufl.inner(u, v) * self._dx(metadata=metadata)
self._lhs = df.fem.form(
a, jit_options=jit_options, form_compiler_options=form_compiler_options
)
self._A = df.fem.petsc.assemble_matrix(self._lhs)
self._A.assemble()
# Create vectors to store right hand side and the solution
self._x = df.fem.Function(space)
self._b = df.fem.Function(space)
# Create Krylov Subspace solver
self._ksp = PETSc.KSP().create(space.mesh.comm)
self._ksp.setOperators(self._A)
# Set PETSc options
prefix = f"projector_{id(self)}"
opts = PETSc.Options()
opts.prefixPush(prefix)
for k, v in petsc_options.items():
opts[k] = v
opts.prefixPop()
self._ksp.setFromOptions()
for opt in opts.getAll().keys():
del opts[opt]
# Set matrix and vector PETSc options
self._A.setOptionsPrefix(prefix)
self._A.setFromOptions()
self._b.x.petsc_vec.setOptionsPrefix(prefix)
self._b.x.petsc_vec.setFromOptions()
def reassemble_lhs(self):
df.fem.petsc.assemble_matrix(self._A, self._lhs)
self._A.assemble()
def assemble_rhs(self, h: ufl.core.expr.Expr):
"""
Assemble the right hand side of the problem
"""
v = ufl.TestFunction(self._b.function_space)
rhs = ufl.inner(h, v) * self._dx
rhs_compiled = df.fem.form(rhs)
self._b.x.array[:] = 0.0
df.fem.petsc.assemble_vector(self._b.x.petsc_vec, rhs_compiled)
self._b.x.petsc_vec.ghostUpdate(
addv=PETSc.InsertMode.ADD_VALUES, mode=PETSc.ScatterMode.REVERSE
)
self._b.x.petsc_vec.ghostUpdate(
addv=PETSc.InsertMode.INSERT_VALUES, mode=PETSc.ScatterMode.FORWARD
)
def project(self, h: ufl.core.expr.Expr) -> df.fem.Function:
"""
Compute projection using a PETSc KSP solver
Args:
assemble_rhs: Re-assemble RHS and re-apply boundary conditions if true
"""
self.assemble_rhs(h)
self._ksp.solve(self._b.x.petsc_vec, self._x.x.petsc_vec)
return self._x
def __del__(self):
self._A.destroy()
self._ksp.destroy()
def initial_conditions(S, V):
u0 = df.fem.Function(S)
u0.interpolate(lambda x: np.zeros(x[: S.mesh.topology.dim].shape))
D0 = df.fem.Function(V)
D0.interpolate(lambda x: np.exp(-(((-x[1] - 1) / 0.1) ** 2)))
return (u0, D0)
def energy(u, D, H, g):
kinetic = 0.5 * H * ufl.dot(u, u) * ufl.dx # 0.5*H*dot(u, u)*dx
potential = 0.5 * g * D * D * ufl.dx # 0.5*g*D*D*dx
return kinetic, potential
use_gmsh = True
if use_gmsh:
import gmsh
order = 1
res = 0.1
gmsh.initialize()
outer_sphere = gmsh.model.occ.addSphere(0, 0, 0, 1)
gmsh.model.occ.synchronize()
boundary = gmsh.model.getBoundary([(3, outer_sphere)], recursive=False)
gmsh.model.addPhysicalGroup(boundary[0][0], [boundary[0][1]], tag=1)
gmsh.option.setNumber("Mesh.CharacteristicLengthFactor", res)
gmsh.model.mesh.generate(2)
gmsh.model.mesh.setOrder(order)
mesh_data = df.io.gmshio.model_to_mesh(gmsh.model, MPI.COMM_WORLD, 0, gdim=3)
mesh = mesh_data.mesh
else:
import meshio
tmp_data = meshio.dolfin.read(
"../../examples/mixed-poisson/hdiv-l2/meshes/sphere_ico6.xml"
)
mesh = df.mesh.create_mesh(
comm=MPI.COMM_WORLD,
cells=tmp_data.cells_dict["triangle"],
x=tmp_data.points,
e=ufl.Mesh(basix.ufl.element("Lagrange", "triangle", 1, shape=(3,))),
)
x = ufl.SpatialCoordinate(mesh)
# mixed functionspace
S = df.fem.functionspace(mesh, ("RT", 1))
V = df.fem.functionspace(mesh, ("DG", 0))
W = ufl.MixedFunctionSpace(*(S, V))
u, D = ufl.TrialFunctions(W)
w, phi = ufl.TestFunctions(W)
# Set global orientation of test and trial vectors
global_orientation = ufl.sign(ufl.dot(x, ufl.CellNormal(mesh)))
u_ = global_orientation * u
w_ = global_orientation * w
# Setting up perp operator
k = global_orientation * ufl.CellNormal(mesh)
# Extract initial conditions
u_n, D_n = initial_conditions(S, V)
u_n_ = global_orientation * u_n
# Extract some parameters for discretization
dt = 0.05
f = x[2]
H = 1.0
g = 1.0
# Implicit midoint scheme discretization in time
u_mid = 0.5 * (u_ + u_n_)
D_mid = 0.5 * (D + D_n)
# n x u_mid
u_perp = ufl.cross(k, u_mid)
# variational form
F = ufl.inner(u_ - u_n_, w_) * ufl.dx
F -= dt * ufl.div(w_) * g * D_mid * ufl.dx
F += dt * f * ufl.inner(u_perp, w_) * ufl.dx
F += ufl.inner(D - D_n, phi) * ufl.dx
F += dt * H * ufl.div(u_mid) * phi * ufl.dx
a, L = ufl.system(F)
# Preassemble matrix (because we can)
A = ufl.extract_blocks(a)
# Define energy functional
kinetic_func, potential_func = energy(u_n, D_n, H, g)
# Setup solution function for current time
u_h = df.fem.Function(S)
D_h = df.fem.Function(V)
# Predefine b (for the sake of reuse of memory)
b = ufl.extract_blocks(L)
# Set-up linear solver (so that we can reuse LU)
petsc_options = {
"ksp_type": "preonly",
"pc_type": "lu",
"pc_factor_mat_solver_type": "mumps",
"ksp_error_if_not_converged": True,
"ksp_monitor": None,
}
problem = petsc.LinearProblem(
A,
b,
bcs=[],
u=[u_h, D_h],
petsc_options=petsc_options,
petsc_options_prefix="mixed_poisson_",
kind="mpi",
)
u_h, D_h = problem.solve()
# Set-up some time related variables
k = 0
t = 0.0
T = 10.0
# Output initial energy
E_k = mesh.comm.allreduce(df.fem.assemble_scalar(df.fem.form(kinetic_func)), op=MPI.SUM)
E_p = mesh.comm.allreduce(
df.fem.assemble_scalar(df.fem.form(potential_func)), op=MPI.SUM
)
print(t, E_k, E_p, E_k + E_p, D_n.x.petsc_vec.min(), D_n.x.petsc_vec.max())
Es = np.array([[t, E_k, E_p, E_k + E_p]])
# Primers
t = 0
k = 0
V_out = df.fem.functionspace(mesh, ("DG", 2, (mesh.geometry.dim,)))
projector = Projector(
V_out,
{
"ksp_error_if_not_converged": True,
"ksp_typ": "preonly",
"pc_type": "lu",
"pc_factor_mat_solver_type": "mumps",
},
)
_x = projector.project(global_orientation * u_h)
_x.name = f"u_use_gmsh={use_gmsh}"
vtx = df.io.VTXWriter(mesh.comm, f"u_h{use_gmsh}.bp", [_x])
# Time Loop
while t < (T - 0.5 * dt):
u_h, D_h = problem.solve()
# Update u_n
u_n.x.array[:] = u_h.x.array
# Update D_n
D_n.x.array[:] = D_h.x.array
# Update time and counter
t = np.round(t + dt, 3)
k += 1
# Output current energy and max/min depth
E_k = mesh.comm.allreduce(
df.fem.assemble_scalar(df.fem.form(kinetic_func)), op=MPI.SUM
)
E_p = mesh.comm.allreduce(
df.fem.assemble_scalar(df.fem.form(potential_func)), op=MPI.SUM
)
Es = np.vstack((Es, [t, E_k, E_p, E_k + E_p]))
_x = projector.project(global_orientation * u_h)
vtx.write(t)
t = Es[:, 0]
E_k = Es[:, 1]
E_p = Es[:, 2]
E_t = Es[:, 3]
import matplotlib.pyplot as plt
fig = plt.figure()
ax = fig.add_subplot()
ax.plot(t, E_t, label="$E_t$", color="purple")
ax.plot(t, E_k, label="$E_k$", color="red")
ax.plot(t, E_p, label="$E_p$", color="blue", linestyle="--", linewidth=0.5)
ax.set_ylim(0.0, 0.20)
ax.set_ylabel("Energy")
ax.set_xlabel("$t$")
ax.legend()
if use_gmsh:
ax.set_title("Energies obtained using gmsh sphere mesh")
else:
ax.set_title("Energies obtained using sphere_ico4")
plt.savefig(f"energy_gmsh_{use_gmsh}.png")
and energies