Hello,
as the title says, I am having issues with interpolating a CompiledExpression in parallel (with mpirun).
The task I want to solve and the problem can be boiled down to the following: I want to create a certain profile on a two-dimensional mesh. To do so, I solve a one-dimensional PDE and want to use the solution of the PDE to define the two-dimensional profile, by repeating the pattern.
Mathematically, this can be expressed as follows: Let u(x) be the solution to the 1D PDE. Then the 2D profile I want to achieve is given by
v(x,y) = \begin{cases} u(x \mod \alpha) \quad &\text{ if } (x \mod \alpha) \leq \beta \\ 0 \quad &\text{ else} \end{cases}
where \alpha >= \beta. Additionally, for the sake of simplicity, let \beta be the length of the 1D interval where the 1D profile is computed.
This means, that the 1D profile should be repeated (starting from x=0) with a distance given by \alpha. I hope the screenshots attached below will make the problem more obvious.
I have the following MWE which illustrates the problem. The code is given by
from fenics import *
mesh = UnitSquareMesh(32, 32)
W = FunctionSpace(mesh, "CG", 1)
def create_profile():
channel_width = 1 / 8
mesh = IntervalMesh(10, 0.0, channel_width)
dx = Measure("dx", mesh)
V = FunctionSpace(mesh, "CG", 1)
u = TrialFunction(V)
v = TestFunction(V)
a = dot(grad(u), grad(v)) * dx
L = Constant(1.0) * v * dx
def bdry(x, on_boundary):
return on_boundary
bcs = DirichletBC(V, Constant(0.0), bdry)
u = Function(V)
solve(a == L, u, bcs)
cpp_code = """
#include <pybind11/pybind11.h>
#include <pybind11/eigen.h>
namespace py = pybind11;
#include <dolfin/function/Expression.h>
#include <dolfin/function/Function.h>
class Profile : public dolfin::Expression
{
public:
Profile() : dolfin::Expression() {}
void eval(Eigen::Ref<Eigen::VectorXd> values, Eigen::Ref<const Eigen::VectorXd> x) const override
{
Eigen::VectorXd dx(2);
dx[0] = std::fmod(x[0], 0.2);
dx[1] = 0.0;
if (dx[0] > 0.1 | dx[0] <= 0.0) {
values[0] = 0.0;
}
else {
p->eval(values, dx);
const double val = values[0];
values[0] = val;
}
}
std::shared_ptr<dolfin::Function> p;
};
PYBIND11_MODULE(SIGNATURE, m)
{
py::class_<Profile, std::shared_ptr<Profile>, dolfin::Expression>
(m, "Profile")
.def(py::init<>())
.def_readwrite("p", &Profile::p);
}
"""
u.set_allow_extrapolation(1)
expr_u_desired = CompiledExpression(
compile_cpp_code(cpp_code).Profile(), degree=1, p=u.cpp_object(), domain=mesh
)
# using LagrangeInterpolator.interpolate does not work either
# u_des = Function(W)
# LagrangeInterpolator.interpolate(u_des, expr_u_desired)
u_des = interpolate(expr_u_desired, W)
return u_des
u_des = create_profile()
with XDMFFile(MPI.comm_world, "u_des.xdmf") as file:
file.parameters["flush_output"] = True
file.parameters["functions_share_mesh"] = False
file.write_checkpoint(u_des, "u_test", 0, XDMFFile.Encoding.HDF5, False)
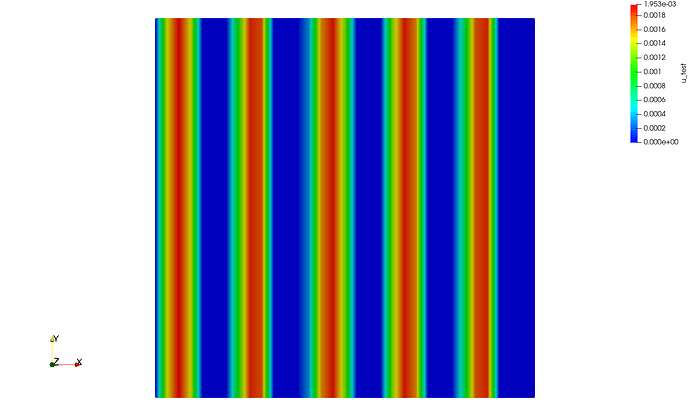
Running this program in serial gives the following output
Which achieves exactly what I am looking for.
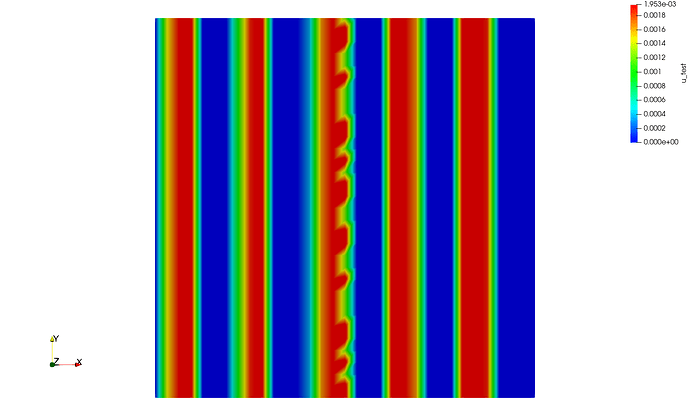
However, when the exact same program in parallel, I get the following results with mpirun -n 2 python script.py
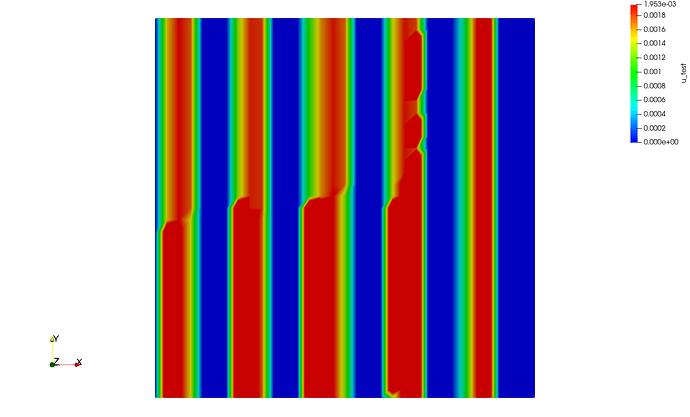
And it only gets worse for 3 MPI processes mpirun -n 3 python script.py
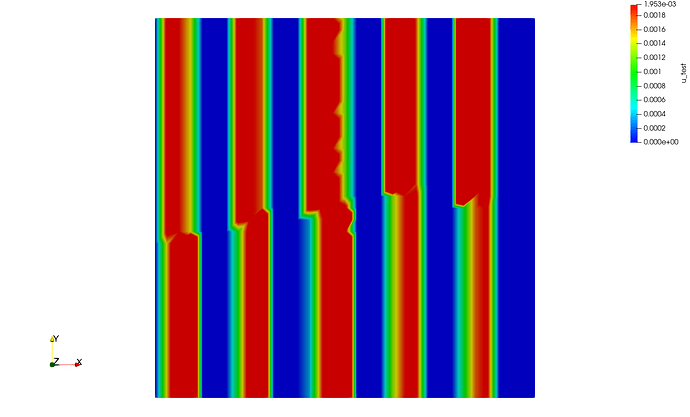
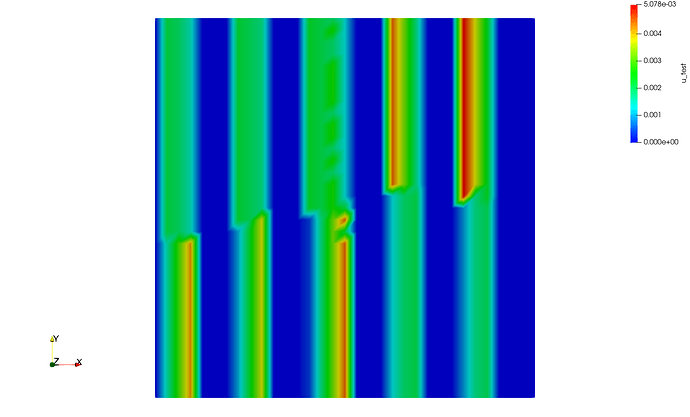
or even 4 processes (mpirun -n 4 python script.py)
and for the sake of completeness, here is the one with 4 processes, but rescaled
As you can guess, this effect is amplified for my real application as I cannot use a regular grid and have to use even more processes. I have not tried using anything but a CompiledExpression, but as the grids I have to use are large, using a UserExpression is not really feasible, which is why I have not tried this.
As you can see in the comment in the source code, I have also tried using LagrangeInterpolator.interpolate, but this gives rise to the same problem.
Ah, and by the way: I am using (legacy) FEniCS, version 2019.1.
Does anyone have a hint how to achieve a consistent interpolation of a CompiledExpression in parallel?